This page will hopefully shed a little light on how I use the raw data files, and interpolate the ice concentration values.
All of the data processing is performed using several FORTRAN routines. The main function of each of these routines will be briefly explained, starting with MAIN.F, and proceeding through the other *.f files in the order in which they are called by MAIN.F.
MAIN.F
The program main.f is the "interactive" part of the interpolation routine. Since all data sets contain monthly values for more than one year, the user must define the number of years to be processed by changing the parameter NYR in the declarations section. Also, the user can specify the FIRST year to be processed by changing the NSKIP variable. The integer variable NSKIP is set to the number of years to be skipped. For example, if you wish to read in the year 1993, of NSIDC data you must skip 1901 to 1992, inclusive. Thus, NSKIP = 92. Note that every data set "begins" at a different year! (ie: SMMR data starts at 1979 and goes to 1997, while GISS AOM data starts at 1950 and goes to 2099)
Also included in MAIN.F is a variable called METHOD. By setting METHOD equal to 1, 2, or 3, the user is specifying interpolation by gaussian, witch of agnessi, or nearest neighbor method, respectively. These methods will be explained later.
MAIN.F calls the following:
Click HERE to see the FORTRAN code for MAIN.F, as used to process NSIDC data.
The purpose of gdata.f is to define the geographic grid coordinates for the ice data, to read in the ice conc/mass data, and to define the land mask. It is necessary to specify the path name of the source ice data, as well as the land mask and grid coordinates if they were contained within a data file. The NSKIP and NYR variables defined in main.f are used in this subroutine to ensure that only a specified number of years are processed.
For GOSTA data, the ice concentration data, as you will recall, are not located in one large file, as is the case with the other data sets. In order that all required data can be read by the computer without repeating the same section of code for each of the 26 data files from 1949 to 1994, an ASCII text file containing the path names of each data file was created. A READ statement was then used to read an individual file name each time a new GOSTA data file was required.
Click HERE to see the FORTRAN code for gdata.f, as used to read GOSTA
All data values, once read into arrays, are normalized such that the ice concentration values range from 0 to 1. Also if ice mass per unit area is used, as in CCCMA and GISS, then the ice mass values are converted to ice concentration values in this subroutine, using the conversion discussed in the previous section. GDATA.F returns an array of ice concentrations, arrays of lat. + lon. values, and a land mask array. Note that for GOSTA, land values are flagged directly in the ice concentration records, so no separate land mask array was necessary.
In this subroutine, which is identical for all data sets, the arctic model grid is defined, and an array containing the arctic land mask is created by reading in a file called "kmt.arc", which contains all of the land mask data. The arctic grid areas are 0.5° by 0.5°, and the dimensions of the grid are 91x67.
Click HERE to see FORTRAN code for RDATA.F
This subroutine performs the interpolation. First, a nested do loop is created which loops through all of the i longitudes and j latitudes of the arctic model grid. Each arctic ocean grid point within this (I,J) loop is then rotated onto the geographic grid which contains the ice concentration values. This rotation is specified by the Eulerian angles located near the top of MAIN.F. The subroutines which perform the rotations were written by Michael Eby from the University of Victoria.
Click HERE to see the FORTRAN code and documentation for ROTATE.F
Once the data is rotated, the arc distance between the rotated grid point and EVERY ice conc. data point, if defined as an ocean point by the mask created in GDATA.F, is calculated. This distance calculation is performed in the FORTRAN routine DIST.F, and this routine is called in INTP.F.
Click HERE to see the FORTRAN code for DIST.F
If either GAUSSIAN or WITCH interpolation was selected in MAIN.F, one of the following two calculations will be performed, to determine a weight for each ice concentration value:
Figure 2.1
The interpolated data value on the rotated arctic grid is determined by dividing the sum of all of the ice concentration values multiplied by their calculated weights, by the sum of all the weights.
intp data val = (sum of (val*wt) / (sum of all wt))
For the GOSTA and NSIDC grids, the value of radius is set to 0.5°. For model output, the value of radius is set to 1.5°. For SMMR, the value of radius is set to 0.2°.
Radius:
The following few paragraphs discuss the reasoning behind choosing the 0.5 radius for the GOSTA and NSIDC data sets. An analogous argument can also be applied to model output.
Since the distance between the rotated arctic grid point and the closest geographic data point will not be greater than 1.414° (sqrt(2)) - the diagonal distance across a 1° cell - it is important that the geographic data with distances greatly exceeding 1.414° are given a sufficiently small weight, such that the total contribution from these distant points do not significantly impact the interpolated data value calculation.
A radius value of 0.5° has been chosen for the NSIDC and GOSTA routines. By increasing the radius value, the interpolated data values will increase, as distant points will have a larger weight. The converse will happen if R were to decrease. Comparing gaussian to witch interpolation, by comparing the average ice conc. value as given in IDL, the results are similar to only one, occasionally two significant figures.
The following seven figures illustrate the difference between interpolation schemes and radii:
[Click on images to enlarge]
Figure 2.2a - Gaussian, R=0.2 Figure 2.2b - Witch,
R=0.2


Figure 2.3a - Gaussian, R=0.5 Figure 2.3b - Witch,
R=0.5


Figure 2.4a - Gaussian, R=1.0 Figure 2.4b - Witch,
R=1.0
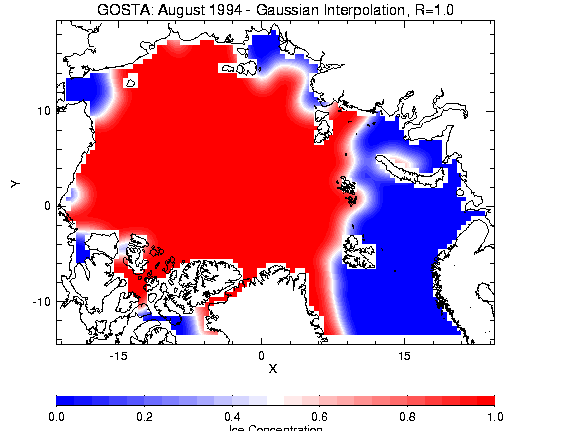

Figure 2.5a - Gaussian, R=2.0 Figure 2.2b - Witch,
R=2.0

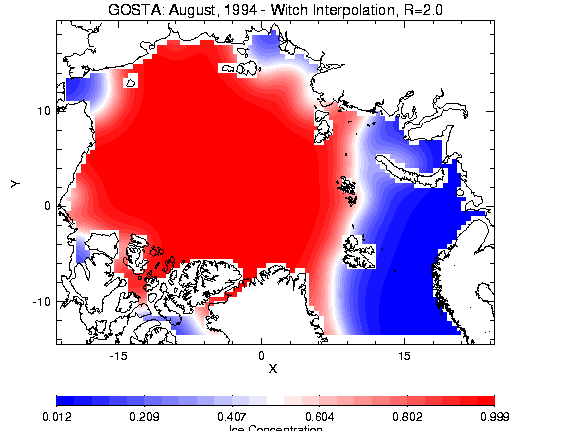
For the SMMR data, which had a grid spacing with finer resolution (~0.2°) than the rotated arctic grid (0.5°), a radius of 0.2° was chosen due to the consideration that within a 0.2° radius of an arctic grid point, several ice concentration points may be present.
If the nearest neighbor method of interpolation is selected in MAIN.F, the rotation and distance calculations, as mentioned above, are calculated. As the program loops through each individual arctic grid point, the geographic point NEAREST to the rotated arctic grid point is found. The corresponding ice concentration value for this nearest geographic point is assigned to the rotated arctic grid point-no weighting occurs. Please see Figure 2.6.
After the interpolation is completed, and the program exits from INTP.F, the array containing all of the interpolated data values is written to an unformatted data file. This procedure is done as a time saving device, since it is only necessary to perform the interpolation once.
Click HERE to see INTP.F as written for the NSIDC data set.
Last Modified: February 2001
Maintained by Erica Muzzerall
[email protected]
Updated by Daniel Roberge
[email protected]
![[JPOD Home]](../../images/orca5.jpg)